For the first time, scientists have been able to sequence the complete human genome, revealing never-before-seen regions of DNA that code for things like muscular dystrophy, some cancers, and even our big brains.
When the human genome project completed its sequencing in 2003, it offered an accurate picture of 92 per cent of our genome. The remaining 8 per cent was too complex for the technology of the time to map, and since then researchers have been working to provide a full picture of the DNA blueprint that makes us human.
The team behind the discovery – named the Telomere-to-Telomere Consortium (T2T) – say the newly mapped areas of our genome could lead to innovative treatments and greater understanding of human variation.
What is the human genome?
The human genome is the entirety of a person's DNA. It includes around 21,000 genes, which, if removed from a single cell and stretched out, would form a strand around 2 metres long. These genes are segmented, and are often found grouped in 46 bundles, which are known as chromosomes.
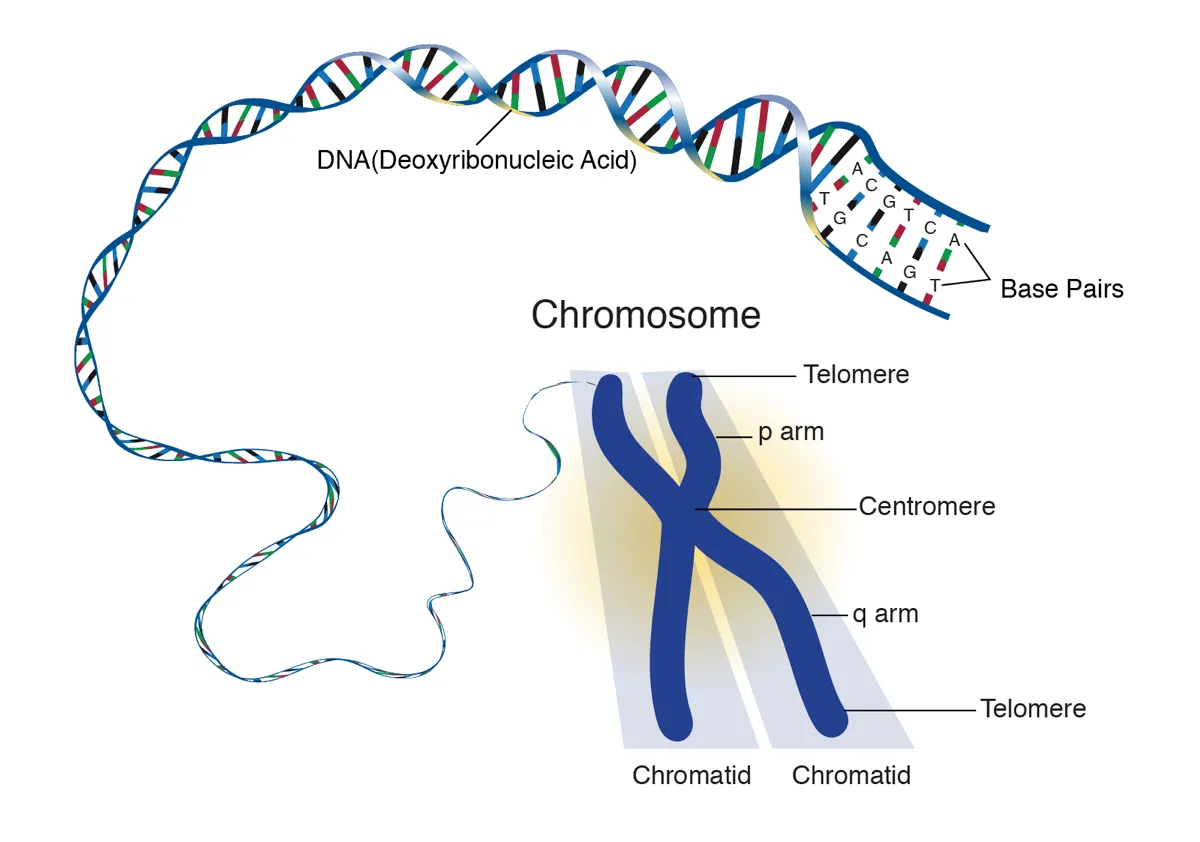
"The genome is the complete book of instructions for a species, and every species has its own genome. It’s basically the blueprint that tells cells how and when to create an individual of that species," said Dr Karen Miga, assistant professor of biomolecular engineering at UC Santa Cruz.
Along with Dr Adam Phillippy of the National Human Genome Research Institute, Miga lead an international team of scientists in the search for the missing sections of the human genome.
"[When the first genome was sequenced in 2003] we had the book, but there were whole chapters missing. Over the years we’ve added pages here and there, unscrambling some text or correcting some errors. But now, we go from page one to the final page of the book. Having this will help us better understand how we form, as an organism, and how we vary among other humans and other species," said Miga.
What was found in our missing DNA?
The 8 per cent hidden represented a huge gap in medical and genomic knowledge. "On a world map, 8 per cent is about the size of Africa, so almost an entire continent was missing," said Miga.
Among the new information sequenced were two crucial areas of our chromosomes: the telomeres and the centromeres.
Telomeres are small caps found at the end of our chromosomes. It is these areas of our chromosomes that shorten as we age. Some research has shown that people who have shorter telomeres than is normal for their age are at increased risk of diseases of old age, and die sooner than people with longer ones. Studies suggest the lengthening of telomeres is one possible way to slow down ageing.
Read more about our DNA:
- Beyond DNA: How proteins let us get up close and personal to our ancient relatives
- A neuroscientist explains how your genes affect your mental health
The centromeres are the structures in the middle of the chromosome. This area is important during cell division, as it's at this site of constriction that proteins attach and split up the DNA so it can be shared between the two new cells.
"Problems or error in the centromeres is something we see in cancers, in ageing and in infertility,” said Miga.
The new genome, called T2T-CHM13, revealed nearly 2,000 candidate genes that researchers can now study further.
Some sections have already been identified by the team to contain genes linked to our immune response, that help us to survive infection and plagues, said Prof Evan Eichler, of the Eichler Lab in the department of genome sciences, University of Washington.They also found important areas that can predict how a person might respond to certain drugs and treatments, said Eichler.
“Some of these [new findings] are the genes that make us uniquely human – half of the genes thought to be responsible for our bigger frontal lobes, making our brain bigger than other primates, are in these newly sequenced regions,” said Eichler.
Once sequenced, the team compared their new human genome to other, existing samples of DNA, and over one million new genetic variants, many of which had never seen before.
"Within previously unresolved regions of the genome, we identified hundredsof thousands of variants... a promising opportunity for evolutionary and biomedical discovery," wrote the researchers.

Genetic variants are already being used to aid treatments for some conditions, for example to target specific types of breast cancers with specialised therapies, and it’s thought that the new genome map will only bring more genetically linked diseases to light.
"We’re already seeing some [genome-specific treatments], but I expect in my young son’s lifetime we’ll see more and more as we learn new things about the role of genetic variants," said Prof Michael Schatz, the co-lead for T2T's variance team.
“We’re finding variants that are only detectable using these new technologies. Are these causal [in relation to diseases]? We can’t say that. But the fact there is an entire class of variants that we haven’t been able to see before, and it’s very exciting to me," said Schatz.
Whose DNA was used in the new human genome project?
The genome used in the research “represents no person that ever lived”, Phillippy said. It came from a special cell created when a sperm and joined an empty egg. Unlike normal cells that come from a union of sperm and egg, which contain half the DNA of the mother and half of the father, this specialised cell contained only the sperm's genetic information. It was collected over 20 years ago, and the donors remain anonymous.
